May 1, 2024
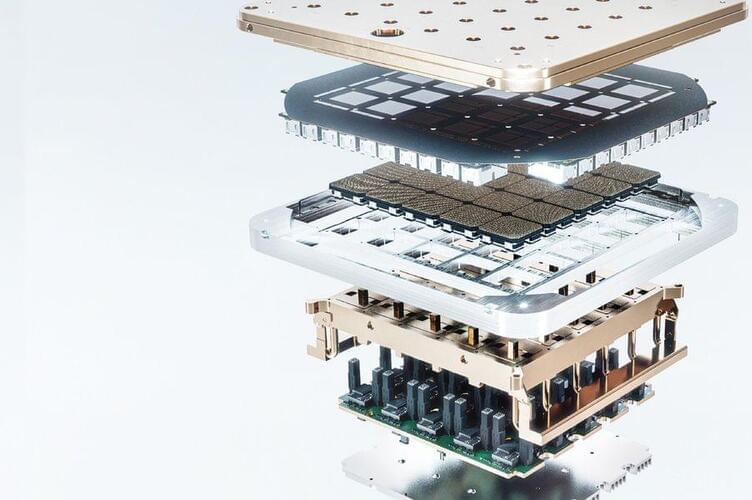
Physicists build new device that is foundation for quantum computing
Posted by Dan Breeden in categories: computing, quantum physics
Scientists have adapted a device called a microwave circulator for use in quantum computers, allowing them for the first time to precisely tune the exact degree of nonreciprocity between a qubit, the fundamental unit of quantum computing, and a microwave-resonant cavity. The ability to precisely tune the degree of nonreciprocity is an important tool to have in quantum information processing. In doing so, the team derived a general and widely applicable theory that simplifies and expands upon older understandings of nonreciprocity so that future work on similar topics can take advantage of the team’s model, even when using different components and platforms.